In a new paper [PDF] appearing in Science, a team of IPD researchers together with colleagues at UW Medicine and Fred Hutchinson Cancer Research Center demonstrate a new way to precisely target cells — including those that look almost exactly like their neighbors. They designed nanoscale devices made of synthetic proteins that target a therapeutic agent only to cells with specific, predetermined combinations of cell surface markers.
These ‘molecular computers,’ which are based on the LOCKR system, operate all on their own, searching out the cells that they were programmed to find.
“We were trying to solve a key problem in medicine, which is how to target specific cells in a complex environment. Unfortunately, most cells lack a single surface marker that is unique to just them. So to improve cell targeting, we created a way to direct almost any biological function to any cell by going after combinations of cell surface markers.”

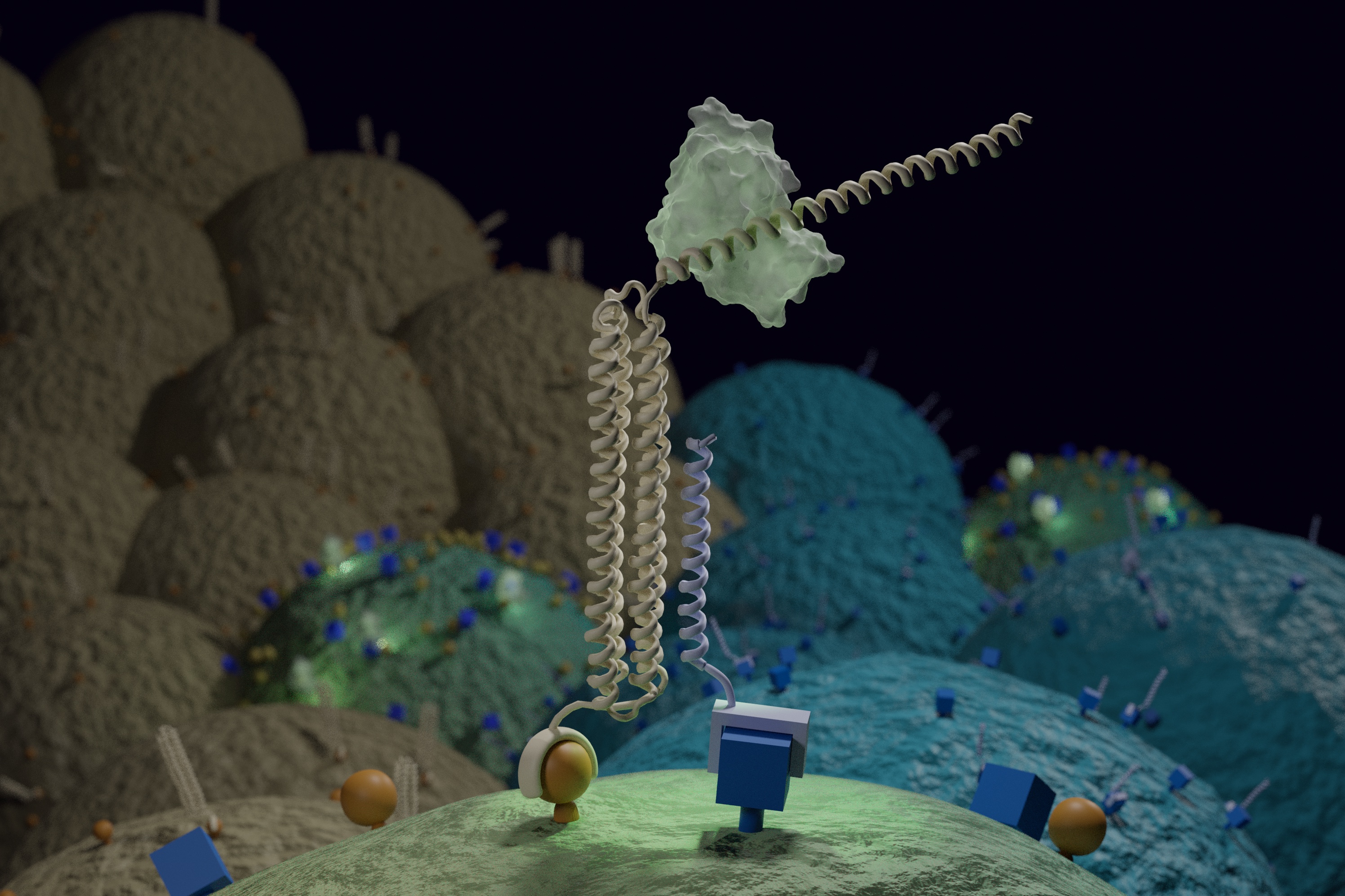
The tool they created is called Co-LOCKR, or Colocalization-dependant Latching Orthogonal Cage/Key pRoteins. It consists of multiple synthetic proteins that, when separated, do nothing. But when the pieces come together on the surface of a targeted cell, they change shape, activating a sort of molecular beacon.
The presence of these beacons on a cell surface can guide a predetermined biological activity — like cell killing — to a specific, targeted cell.
The team showed that Co-LOCKR can focus the cell-killing activity of CAR T cells. In the lab, they mixed Co-LOCKR proteins, CAR T cells, and a soup of potential target cells — some had just one marker, others had two or three. Only the cells with the predetermined marker combination were killed by the T cells. If a cell also had a predetermined “healthy marker”, that cell would be spared.
“T cells are extremely efficient killers, so the fact that we can limit their activity on cells with the wrong combination of antigens yet still rapidly eliminate cells with the correct combination is game-changing.”

This cell-targeting strategy relies entirely on proteins, which sets it apart from most other methods that rely on engineered cells and operate on slower timescales.
This research was conducted at the University of Washington School of Medicine Institute for Protein Design, the Immunotherapy Integrated Research Center at Fred Hutchinson Cancer Research Center, and the University of Washington Department of Bioengineering.
The co-lead authors of this work are Marc J. Lajoie (supported by a Washington Research Foundation Innovation Postdoctoral Fellowship and a Cancer Research Institute Irvington Fellowship from the Cancer Research Institute), Scott E. Boyken (supported by the Burroughs Wellcome Fund Career Award at the Scientific Interface), and Alexander I. Salter (supported by the Hearst Foundation and Fred Hutchinson Cancer Research Center Interdisciplinary Training Grant in Cancer Research). This work was also supported by the National Institutes of Health (R01 CA114536, NIGMS T32GM008268, 1R21CA232430-01, T32CA080416), the National Science Foundation (CHE-1629214), the Defense Threat Reduction Agency (HDTRA1-18-1-0001), the Nordstrom Barrier IPD Directors Fund, the Hearst Foundation, the Washington Research Foundation and Translational Research Fund, the Howard Hughes Medical Institute, the Open Philanthropy Project, and The Audacious Project organized by TED.